Big Tree: Instructions
The Big Tree project is open to all men who wish to contribute their NGS results. It's free. This project is one of many that relies on the open sharing of BigY, FGC, or other results. These tests are useful only when the data is compared to those of other men.
Due to the recent shutdown of the ydna data warehouse in May, 2025, you must contact me directly at (scotsgenealogy at gmail.com) to add your data to this project.
Big Y
For the BigY, the files I need are the VCF files. This consists of a .BED and .VCF file which can be downloaded from the BigY results page for free. They come as one .zip file, which is about 10MB or so in size. Please don't extract the files from the .ZIP archive, leave them as they come.
The surname that appears in the filename is the surname I'll use with your kit number on the tree. You can always e-mail me to make changes.
Your "Raw Data" is accessed from your FTDNA page. After you sign in click on the link for your "Big Y Results".
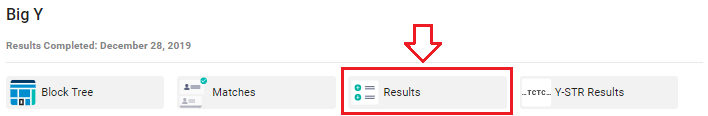
There is a blue "Download Raw Data" button in the upper right corner of the window.
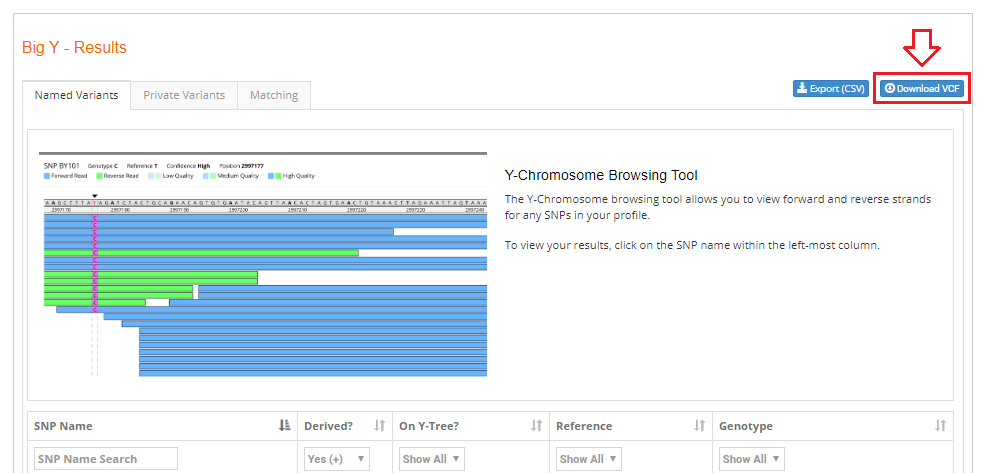
Finally click the green "Download VCF" button in the windows the opens.

This will prompt you to save the .zip file to your computer. Please don't modify this file before uploading it.
To upload to the Y-DNA Data Warehouse, you are better to provide a link to the VCF file, rather than trying to upload directly.
Y-DNA Data Warehouse Instructions
Big Y BAM File
The BAM file for BigY results is accessed in much the same way as the raw, VCF data. By first contacting FTDNA via https://www.familytreedna.com/contact.aspx#contactForm you can request access to the BAM file for your results. This file is the true raw data for the BigY test. It is a large file, generally about 1GB or so in size, and you need special software to read it. I generally don't need your BAM file, but it is useful to refer to for some situations. If you have it available, please forward the link to me. If you are submitting your BigY data to FullGenomes or YFull for further analysis, they will also need access to your BAM file.
FGC
The FGC "Interpretation Results" can be shared in a number of ways. Just as for the BigY results above, data should be uploaded to the Y-DNA Data Warehouse. To download the results, the instructions are:
- Log into your FGC account.
- Click on "My Profile" at the top if you are not already on that page.
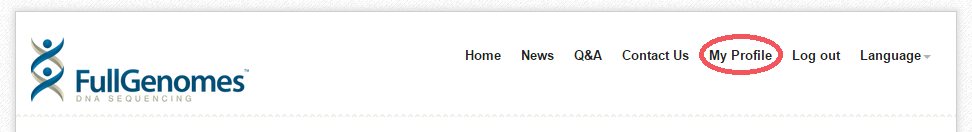
- Under the Results heading, find your name, and below that click the "Interpretation Results" link.
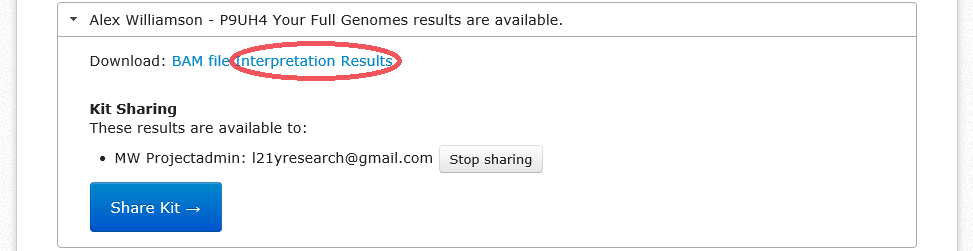
If your haplogroup is R-L21, you can share your results with the R-L21 researchers from within the FGC website by sharing them with "l21yresearch@gmail.com". If you're not R-L21, you can still share your results directly with me by using "scotsgenealogy@gmail.com". This is useful for giving me access to the BAM file for your FGC results. Even if you "share" the results this way, you still need to upload them to the Y-DNA Data Warehouse.
- Instead of clicking on "Interpretation Results", click the blue button labelled "Share Kit".
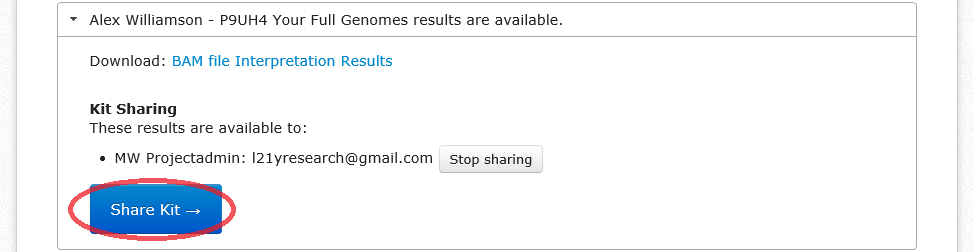
- A new window will open. Enter the e-mail address "l21yresearch@gmail.com" in the box.
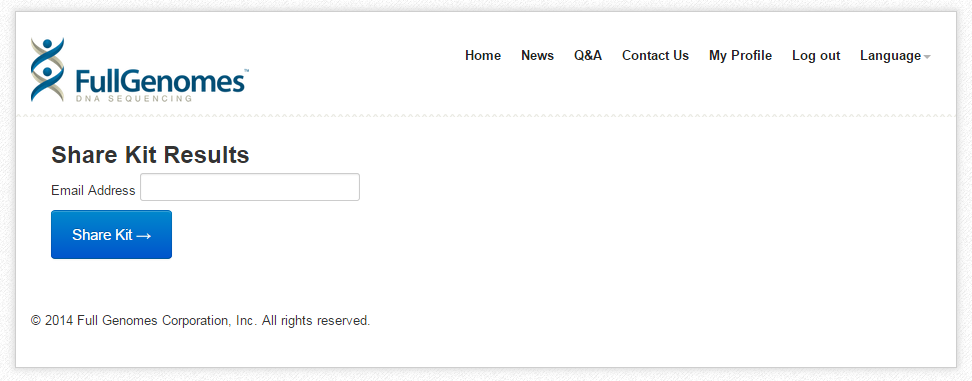
- Click the blue "Share Kit" button.
Haplotype Data
I have started to collect haplotype data for those men on the tree, but I have not started to use it yet. For now, I am only interested in the FTDNA Y-STR results, and not any that can be extracted from NGS tests. I intend to occasionaly download the STR data from the major FTDNA haplogroup projects. Should you decide to e-mail me your Y-STR results, please send them to me as plain text as if copy and pasted from a FTDNA results webpage. For example, my own would be: